BUCKLER LAB
HYBRID VIGOR
The exponential growth of the world's human population is leading to increased food demand, which raises food costs, diminishes arable land, and reduces available water.
Today, maize is the largest production crop in the world. According to the Food and Agriculture Organization of the UN, in 2018, just over 1.4 billion Mg of maize was produced worldwide, compared to 996 million Mg of paddy rice (Oryza sativa L.), and 865 million Mg of wheat (Triticum aestivum L.) . Given the profound impact maize has on the world, any additional knowledge we can acquire to maximize production is highly desirable.

Maize growing in Malawi
The development of hybrid cultivars was an enormous breakthrough in the breeding of maize over time. Maize hybrids were first commercially grown in the 1930s. At the time, U.S. production of maize was 53 million Mg annually. By the 1950s maize production grew to 76 million Mg. It was up to 150 million Mg annually in the 1970s and 219 million Mg in the 1990s (USDA-National Agricultural Statistics Service). This tremendous increase in yield is a combination of improved agronomic management, such as fertilizers, as well as breeding efforts, such as more uniform and higher yielding hybrids that are tolerant to higher density planting.
Understanding Heterosis Key to Increasing Yields
In the U.S., the increase in maize yield over time resulted in the reduction of necessary agricultural land by approximately 13 million hectares (ha) and allowed 98% of Americans to pursue non-agricultural occupations. Today maize is grown on 34 million ha in the U.S., and is a base commodity crop supporting the $1-2 trillion U.S. food industry by providing protein, oil, and starch for food, animal feed, ethanol, and other bio-based products. The only way to sustainably increase yield without increasing inputs or area under cultivation is to continually improve the genetic adaptation of maize to modern farm management and changing climate conditions. Understanding the genomic characteristics underlying heterosis will be critical to increasing maize yield in the U.S. and the world in a sustainable way. In addition to maize, understanding heterosis is also very important in other plant and animal species since hybrids do not only exhibit increased yields but are also more tolerant compared to their inbred parents.
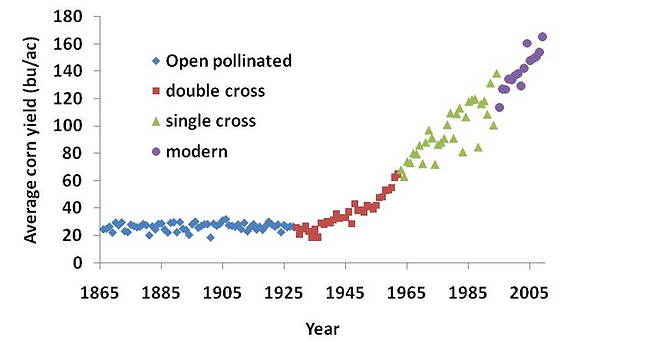
Average corn yield in bushels per acres in the U.S. between 1865 and 2005. (USDA-NASS)
Inbreeding Depression
Inbreeding depression and heterosis have been studied for decades. Inbreeding depression is most commonly observed when outcrossing species, such as maize, are exposed to inbreeding. The progressive selfing or sibling matings reduce the level of genome-wide heterozygosity and thereby the overall fitness of the organism in the form of reduced growth rate, viability, developmental stability, and fertility. Heterosis, on the other hand, is the superior performing heterozygote progenies as a result of crosses made between different populations. Heterosis was first defined by Shull as `the superiority of heterozygous genotypes with respect to one or more characters in comparison with the corresponding homozygotes (Shull, 1911).’ Since its inception in 1909, the pure line hybrid concept developed by Shull has been extensively used in maize breeding (Shull, 1909). Given that maize is a natural outcrossing species, plant breeders have developed an "unnatural" state of inbred maize that cannot be found in the wild. This may be why maize hybrids express a relatively high heterotic effect.
Not All Hybrids Express Heterosis
All hybrids do not express heterosis. It is believed that the heterotic effect of a hybrid is based on the genetic distance between the two parents and the level of adaptation of the parents. These adaptations include altered flowering time, disease resistance, growth habits, and stress tolerance. It is possible that different sets of genes have been selected for the same trait in different inbred lines resulting in a higher number of active genes in the heterozygote hybrid compared to the homozygous inbred. However, there appears to be a limit to the level of diversity desirable in the construction of a hybrid. A simple possible explanation is that tropical lines are not adapted to temperate conditions and vice versa; so, alleles contributed from one of the parents, depending on the growing location of the hybrid, will not positively impact the fitness of the hybrid.
Heterotic Variation
Variation in heterotic effect among hybrids was first observed in inter-varietal crosses where higher performance was seen in hybrids constructed from parents with different types of endosperm (Richey, 1922). This theory is supported by studies showing that maize plants have the ability to more precisely organize unrelated genomes without modification than closely related genomes in the hybrid nucleus. These results are contradictory to what one might have expected. However, since maize is a natural outcrossing species, it may have developed such mechanisms to prevent inbreeding. If this is true, the best performing, most stable hybrids would be those with the most stable genome organization as reflected by normal DNA inheritance (Biradar & Rayburn, 1993).
These findings suggest that crosses should be made between geographically or genetically distant parents, which lead to the creation of the first heterotic groups. It is believed that the populations grouped together in different heterotic groups have divergent genetic backgrounds with unique diversity that has diverged due to founder effect, genetic drift, selection, and as a result of post-divergent mutations.
Five Modes of Action Many Attribute to Heterotic Phenomenon
Over the past one hundred years, several theories have been proposed to explain heterosis. However, there remain five primary modes of action which many attribute to heterotic phenomenon. These include dominance, overdominance, pseudo-overdominance, dosage balance, epistasis, and more recently epigenetics. Our research has supported the prevalence of dominance (and associated pseudo-over dominance) and dosage balance as being key.

Hybrids vs. inbreds (Springer & Stupar, 2007)
Maize HapMap
The recent publication of the complete maize reference genome sequence, the first generation maize HapMap, and the fast-evolving high throughput sequencing technologies have given researchers the opportunity to examine recombination rate and divergence from a genome-wide perspective that had not been possible in the past. Maize shows, as do many other species, evidence of relationships between recombination and physical distance. Near the ends of the chromosomes and telomeres, recombination is enhanced, with the opposite phenomenon holding true in the centromeric regions.
Heterozygosity
A study using the U.S. Nested Association Mapping (NAM) population identified considerable residual heterozygosity in peri-centromeric regions of the maize genome and posited that this retention was probably a consequence of heterosis (McMullen et al., 2009). Regions of increased residual heterozygosity had more than 30% of all genes and nearly average diversity (Gore et al., 2009). Previous studies revealed that anchoring recombination to the physical genome and controlling for chromosome effects, residual heterozygosity and the recombination rate were inversely related, with a correlation of 35%. Conversely, gene density and diversity were much less related to residual heterozygosity. The strong correlation between recombination rate and the level of residual heterozygosity result when controlling for recombination rate, gene density and diversity have a non-significant effect on residual heterozygosity (Gore et al., 2009).
Recombination and Heterozygosity
The relationship found between recombination and heterozygosity suggests that recombination rate is the major factor maintaining residual heterozygosity. Suppressed recombination in these genetically divergent centromeric regions hinders the optimum allelic combination to be formed. This evidence argues for pseudo-overdominance to be causing heterotic effect through linkage of dominant loci acting in repulsion. These regions of suppressed recombination seem to have affected the effectiveness of selection during development of maize inbreds, since this selection has had to work against a genomic region with very strong recombination suppression, and are likely to play a major part in heterosis. Prior studies of the NAM population and analysis of genetic diversity characterized in the first generation maize HapMap suggest that there is increased residual heterozygosity in pancentromeric regions of the maize genome. We infer a direct relationship between recombination rate and residual heterozygosity, which we will examine further.

Every chromosome has more residual heterozygosity in the
recombination suppresed centromeres. (McMullen et al., 2009)

Recombination predicts residual heterozygosity. (Gore et al., 2009)
Dominance effects and the origin of genetic effects in maize hybrids
We utilized maize hybrids created from the North Central Regional Plant Introduction Station (Ames) population to determine the role of genetic dominance (complementation effects at individual loci). We analyzed different types of dominance effects in grain yield, plant height, and flowering time: polygenic effects, QTL effects from GWAS, and directional dominance from analysis of inbreeding depression. We found consistent evidence for dominance effects for plant height and grain yield. For flowering time, genetic effects at play could be more complex. In addition to dominance, there could be substantial epistasis, i.e., complementation effects involving multiple loci or genes, acting in signaling pathways, protein complexes, etc. (Ramstein et al., 2020).
Based on maize hybrids created from the Ames and NAM populations, we showed that variability in maize hybrids arise mostly from loci close to genes (within 1 kb of gene models). Combined with our results about dominance, our results on Ames and NAM hybrids suggest simple complementation effects in genic regions for grain yield and plant height. Furthermore, we showed that models based on enrichment of genetic effects in genic regions, and other functional annotations like evolutionary constraint at loci, could improve genomic prediction across populations (Ramstein et al., 2020). Following this framework, where genomic models are trained in one population and validated in a different panel, we are currently testing methodologies to prioritize loci at a finer scale and detect causal variants within gene regions.